STR loci are defined regions of DNA containing multiple copies
of short sequences of bases (typically 3 to 7 bases) which are
repeated a number of times; the number of repeats varying among
individuals in the population.
| Reference in Figure |
Actual Sample Code |
| Clear (control) |
NHBZ0/0005 |
| Red (spCJD-1) |
NHBX0/0001 |
| Green (spCJD-2) |
NHBX0/0002 |
| Blue(nvCJD) |
NHBY0/0003 |
| Kn52 |
Promega Control Sample |
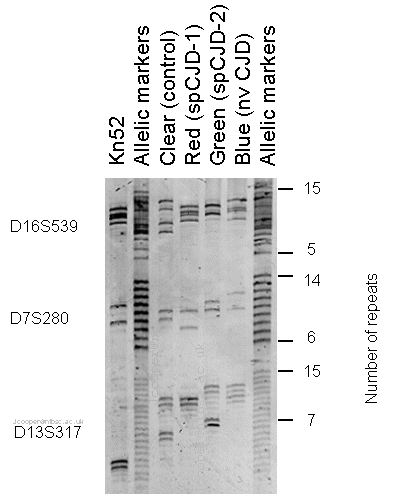
Note: Due to the banding pattern the
samples are distinct (different number of repeats at each loci)
The following data represents DNA
fingerprints of STR (short tandem repeat) loci for our reference
materials. This is to ensure that reference materials are from a
species specific source and are distinct.
STR loci are among the most informative polymorphic markers in the
genome.
The STR analysis procedure used at the CJD resource Centre involves
the simultaneous amplification of eight STR loci and the amelogenin
gene in a multiplex PCR reaction (Promega PowerPlex® 1.2 system,
Promega Corporation). This allows for discrimination of fewer than
1 in 108 individuals (Lins, A.M., et al. JOFS. 43: 1168,
1998). The amplicons are separated on an ABI 310 Prism Capillary
Genetic Analyser and data collected using Genescanning® software
(Applied Biosystems).
Each peak in the resulting electropherogram represents an allele,
is compared to an allelic ladder profile.
Comparative alignment of traces (and manually read repeat numbers)
have shown the samples are distinct and therefore different.
Please Note: The CJD Resource Centre does not
endorse, warranty the use of, or represent the products or
processes used to obtain the following data for any purpose. The
data below is provided on a 'as is' basis and we advise recepients
of samples who demand a specific DNA profile to recharacterise the
samples.
Please note: Amelogenin is not a STR loci and the
data below just indicates size of the band.
Sample: NHBZ00005 -
Clear
This data is no longer available on
this site.
However the samples are distinct and free from cross
contamination.